A truly systems view of life must account for the remarkable property that cells can maintain and reconstitute themselves despite a constant turnover of chemical components. Much research done in systems biology rarely tackles this thorny dynamical problem directly, largely focusing on modeling of individual elements or pathways, or examining genome-wide static patterns. Partly this is due to the sheer complexity of even the simplest cell, partly because the data is still sparse, and partly because many of the theoretical constructs are not easily accessible or intuitive to most biologists.
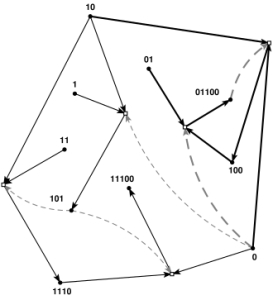 “Autocatalytic sets” – originally introduced several decades ago by Stuart Kauffman – are one of several systems-level models developed to explain the self-maintenance phenomenon and have also been used as possible models for the origin of life. These concepts and models but have been extended considerably in the intervening decades by various scientists including Mike Steele and Wim Hordijk. I have also done some work along these lines with Walter Fontana on “algorithmic chemistry“. In a recent blog post, Hordijk defines an autocatalytic set as a set of chemical reactions in which:
“Autocatalytic sets” – originally introduced several decades ago by Stuart Kauffman – are one of several systems-level models developed to explain the self-maintenance phenomenon and have also been used as possible models for the origin of life. These concepts and models but have been extended considerably in the intervening decades by various scientists including Mike Steele and Wim Hordijk. I have also done some work along these lines with Walter Fontana on “algorithmic chemistry“. In a recent blog post, Hordijk defines an autocatalytic set as a set of chemical reactions in which:
- each reaction in the set is catalyzed by at least one of the molecules from the set itself, and
- each molecule in the set can be produced from a basic food source through a series of reactions from the set itself.
The food source is a subset of molecule types which are assumed to be directly available from the environment …. The above definition captures the idea of life as a self-regulating (part 1) and self-sustaining (part 2) chemical reaction network.
Computer simulations demonstrating the plausibility that the set of chemical reactions within cells constitute a collectively autocatalytic process have been developed over the years, but empirical evidence has been sparse. Hordijk and colleagues recently examined all the chemical reaction networks inside the bacteria E. coli, and integrated it with their existing mathematical and computational framework. In a recent paper in the Journal of Systems Chemistry – described in an associated blog post – they found that 98% of reactions in E. coli appear to form an autocatalytic set. They also found that the set was resilient to environmental and other changes, such robustness being a hallmark of any living organism:
Moreover, the autocatalytic set in E. coli‘s metabolism is quite robust against environmental variations, such as different food sets or the removal of random reactions or molecules. Using our formal framework, we can easily check how important each individual molecule or reaction is to the existence and sustainability of the original autocatalytic set. For example, we can remove a particular molecule or reaction from the metabolic network, and then check whether it still contains an autocatalytic set, and if so, how much smaller it is than the original set.
These are intriguing results, especially with the growth of interest in the human microbiome – the collection of bacterial “flora” within the human body – of which E. coli is but one member. Of course more research needs to be done, but they open up a tantalizing possibility that autocatalytic sets may extend to include chemicals secreted by between different species of flora within the human gut.
Read more…. at “Autocatalytic Sets in Your Gut” on Wim Hordijk’s blog.

Interesting post, but could you clarify why the set of all reactions in a cell is not automatically an autocatalytic set?
LikeLike
It’s a good question. Those reactions that aren’t in the autocatalytic set could be thought of as “side reactions” draining resources away from the autocatalytic cycle. So it does seem intuitive that most (if not all) reactions in the cell would be collectively autocatalytic (otherwise what are they doing there). This is, indeed, what they found (at least in 98% of the E. coli reactions were in the autocatalytic set). But in a sufficiently complicated reaction network, it may be inevitable that there will always be some side reactions present (2% in E. coli), and my sense is that there would be evolutionary pressure to remove those reactions over time. Wim probably has an even better explanation.
LikeLike
My molecular biologist collaborators on this work are the experts on actual E. coli metabolism, so they may have a better idea of what those 2% of reactions are (or do) that are not part of the autocatalytic set. My guess is that it probably also has to do with missing or incomplete data. We took E. coli, because its metabolic network has been studied the most extensively of all living organisms, and is the most complete. But even then, there are still many missing bits and pieces. So maybe we should redo the analysis in another couple of years, when some of that missing data has been filled in.
Another issue is that we did not take “all reactions in the cell”, as you mention. For example, we took that part of the metabolic reaction network that takes place in the cell’s cytoplasm (“interior”), which is the majority of the reactions. But there are also reactions taking place in the periplasm (“boundary layer”), and there are transport reactions, etc. And if we were to look at the entire cell, we’d also have to take into account the genetic system. So it’s only a first step, but in my opinion a very promising (and important) one. As far as we know it’s the first formal proof that a living system (or an essential part thereof) is indeed an autocatalytic set.
Hope that helps — Wim
LikeLike