With warm New England weather finally kicking in, it was nice to get a Spring surprise in the form of our new paper on prions being released. Prions are a particular kind of protein that propagate by imbuing an altered shape, or “confirmation”, on other proteins of the same type. They essentially act as a kind of protein “zombie”, and there has been some speculation that they may support a kind of “memory function” due to their ability to transmit state across generations. Prions were first discovered in the context of diseases like scrapie and variant Creutzfeldt–Jakob Disease (vCJD) but have shown up in all kinds of unexpected places, such as yeast and possibly – providing the aforementioned Spring surprise – plants. Scientific American has a nice blog post on a recently-published study from the Whitehead Institute, authored by Sohini Chakrabortee, Can Kayatekin, Greg Newby, Marc Mendillo and Susan Lindquist, in which I contributed computational analysis. (Apologies for the paywall, I can get you a copy if you contact me).
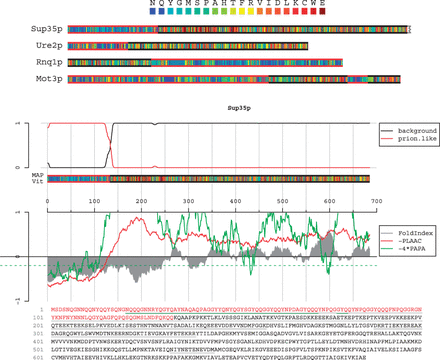 This study is a good example of the power of using genome-wide algorithmic predictions to help prioritize what would otherwise be a needle-in-a-haystack search. In the study, we used the open-source PLAAC web-based tool (stands for Prion-Like Amino Acid Composition) that Oliver King, Andy Nutter-Upham and I developed a few years back (code is available on GitHub, see also Projects and Software) to generate a list of potential candidate prions in the mustard plant, Arabidopsis thaliana from the proteome sequence alone. These candidates were then investigated using biochemical and molecular biology tools to see if they looked particularly prion-y. One interesting protein, LD (or Luminidependens) showed up having strong prion-like properties. LD is also involved in the biochemical pathway for the decision for a plant to flower in Spring after the extended cold of a typical winter (our last 2015-2016 winter here in New England almost didn’t qualify it was so mild). Although it’s way too early to say yet, it’s possible that LD could provide the basis for plants “remembering” this cold state. As the blog post states:
This study is a good example of the power of using genome-wide algorithmic predictions to help prioritize what would otherwise be a needle-in-a-haystack search. In the study, we used the open-source PLAAC web-based tool (stands for Prion-Like Amino Acid Composition) that Oliver King, Andy Nutter-Upham and I developed a few years back (code is available on GitHub, see also Projects and Software) to generate a list of potential candidate prions in the mustard plant, Arabidopsis thaliana from the proteome sequence alone. These candidates were then investigated using biochemical and molecular biology tools to see if they looked particularly prion-y. One interesting protein, LD (or Luminidependens) showed up having strong prion-like properties. LD is also involved in the biochemical pathway for the decision for a plant to flower in Spring after the extended cold of a typical winter (our last 2015-2016 winter here in New England almost didn’t qualify it was so mild). Although it’s way too early to say yet, it’s possible that LD could provide the basis for plants “remembering” this cold state. As the blog post states:
This does not mean that plants definitely have prion-like proteins, adds Lindquist—but she thinks that it is likely. “I’d be surprised if they weren’t there,” she says. To prove it, researchers would need to grind up a plant and see whether they could find a protein such as LD in several different folded states, as well as show that any potential prion caused a misfolding cascade when added to a test-tube of protein. Lindquist adds that because she’s not a plant scientist—her focus is on using yeast to investigate prions—she hasn’t tried these experiments.
PLAAC can also be used on any proteome of interest (you simply need a FASTA file), the code is open-source so for all budding prion-hunters, more intriguing features may await discovery in other organisms…
